# Load libraries
library(data.table)
library(devtools)
library(presto)
library(glmGamPoi)
library(sctransform)
library(Seurat)
library(tidyverse)
library(miQC)
library(SeuratWrappers)
library(flexmix)
library(SingleCellExperiment)
library(SummarizedExperiment)
library(readxl)
library(fishpond)
library(Matrix)
library(speckle)
library(scater)
library(patchwork)
library(vctrs)
library(alevinQC)
library(harmony)
library(scDblFinder)
library(cellXY)
# Set global options for Seurat v5 objects
options(Seurat.object.assay.version = 'v5')6 Skin: Sub-clustering on T/NK lineage
6.1 Set up Seurat workspace
6.2 Load previously saved object
merged.18279.skin.singlets <- readRDS("Skin_scRNA_Part3.rds")6.3 Set idents to preferred initial clustering resolution
Idents(merged.18279.skin.singlets) <- merged.18279.skin.singlets$RNA_snn_res.0.8
merged.18279.skin.singlets$seurat_clusters <- merged.18279.skin.singlets$RNA_snn_res.0.8
DimPlot(merged.18279.skin.singlets, reduction="umap.harmony", label=T)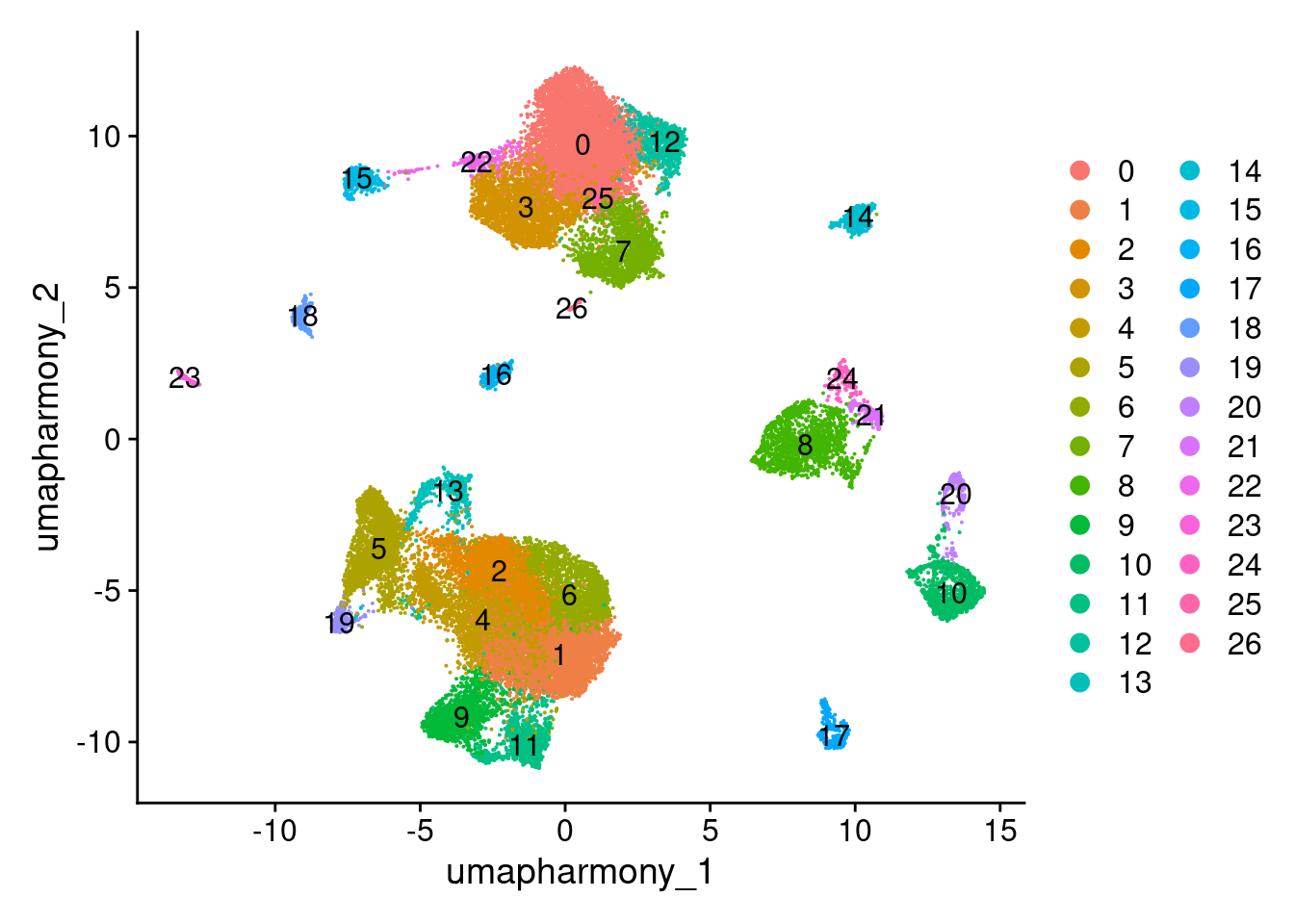
6.4 Run sub-clustering on T/NK lineage
T/NK cells are in clusters 1, 2, 4, 5, 6, 9, 11, 13, 19
6.4.1 Cluster 1
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "1",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 3688
Number of edges: 114356
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8361
Number of communities: 2
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)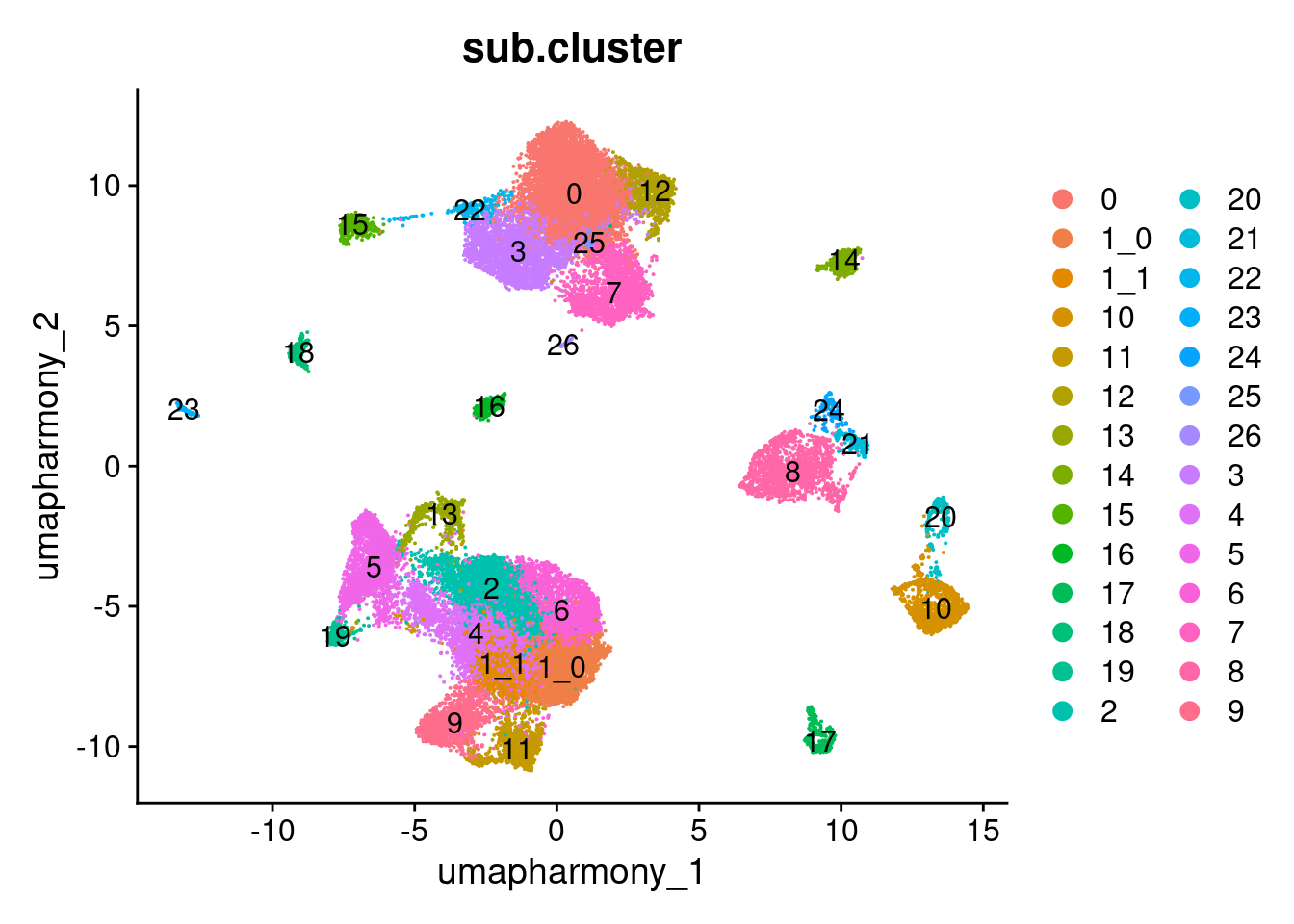
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11 12 13 14 15 16 17 18 19 2 20 21
5768 2588 1100 1123 1086 907 569 557 472 407 360 308 280 2851 262 238
22 23 24 25 26 3 4 5 6 7 8 9
228 177 175 46 33 2418 2363 2200 2186 1746 1483 1466 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13" "14" "17"
[13] "23" "1_0" "12" "4" "11" "9" "5" "6" "2" "16" "19" "22"
[25] "24" "15" "20" "26" 6.4.2 Cluster 2
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "2",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 2851
Number of edges: 89069
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8349
Number of communities: 3
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)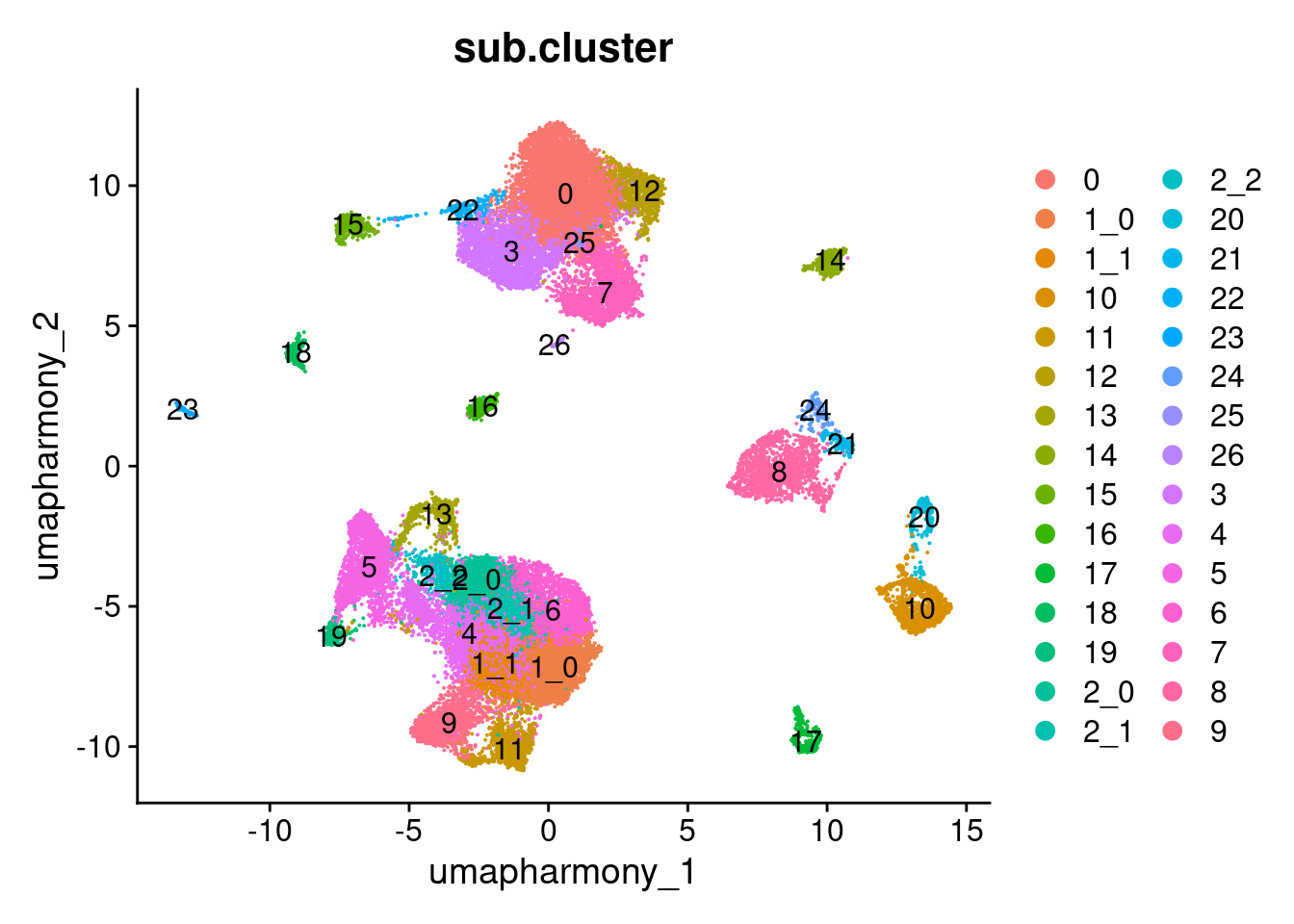
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11 12 13 14 15 16 17 18 19 2_0 2_1 2_2
5768 2588 1100 1123 1086 907 569 557 472 407 360 308 280 1464 980 407
20 21 22 23 24 25 26 3 4 5 6 7 8 9
262 238 228 177 175 46 33 2418 2363 2200 2186 1746 1483 1466 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13" "14" "17"
[13] "23" "1_0" "12" "4" "11" "9" "5" "6" "2_1" "2_0" "16" "19"
[25] "2_2" "22" "24" "15" "20" "26" 6.4.3 Cluster 4
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "4",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 2363
Number of edges: 65111
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8414
Number of communities: 2
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)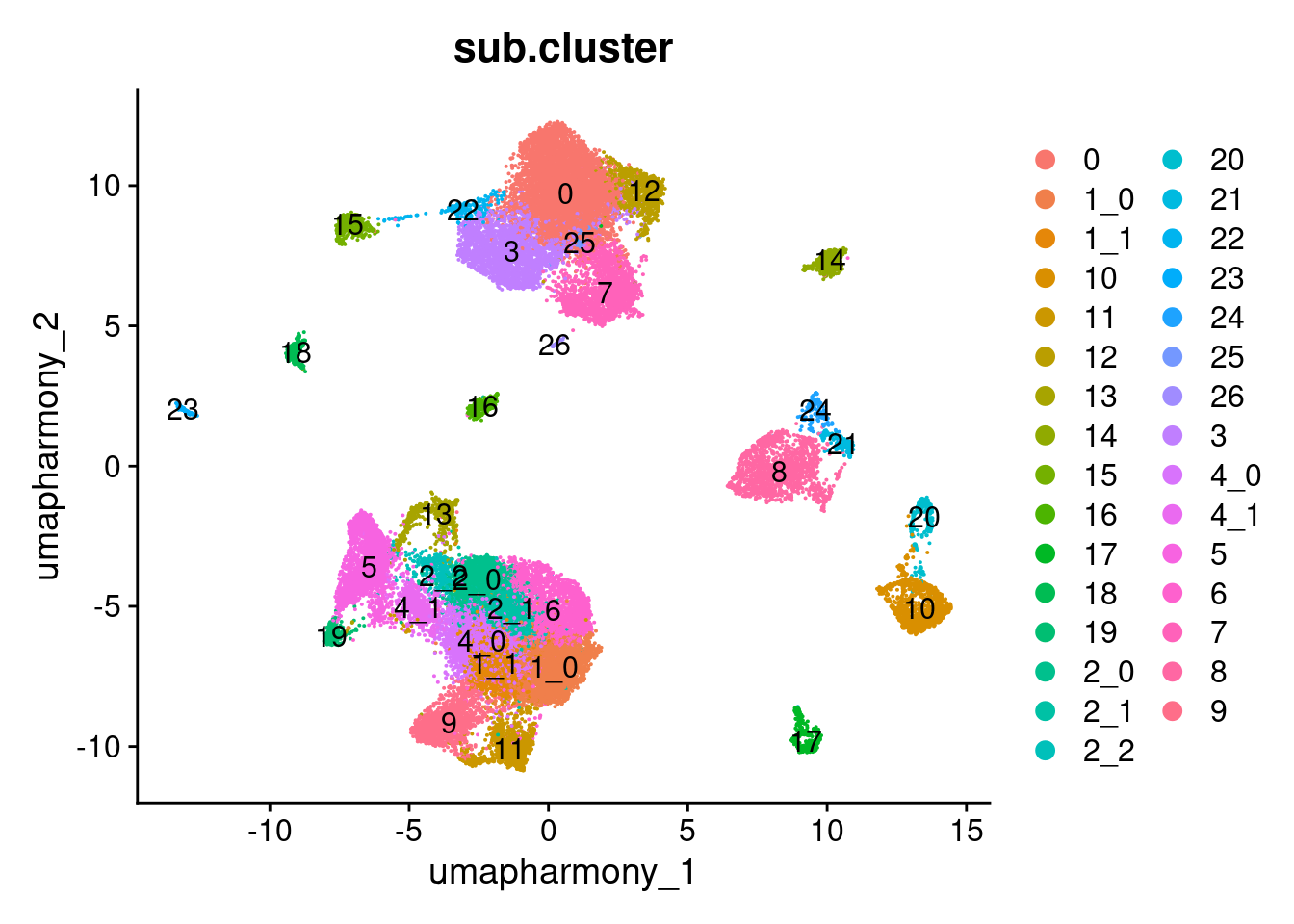
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11 12 13 14 15 16 17 18 19 2_0 2_1 2_2
5768 2588 1100 1123 1086 907 569 557 472 407 360 308 280 1464 980 407
20 21 22 23 24 25 26 3 4_0 4_1 5 6 7 8 9
262 238 228 177 175 46 33 2418 1826 537 2200 2186 1746 1483 1466 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13" "14" "17"
[13] "23" "1_0" "12" "4_0" "11" "9" "5" "6" "2_1" "2_0" "16" "19"
[25] "2_2" "22" "24" "15" "20" "4_1" "26" 6.4.4 Cluster 5
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "5",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 2200
Number of edges: 82669
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8513
Number of communities: 3
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)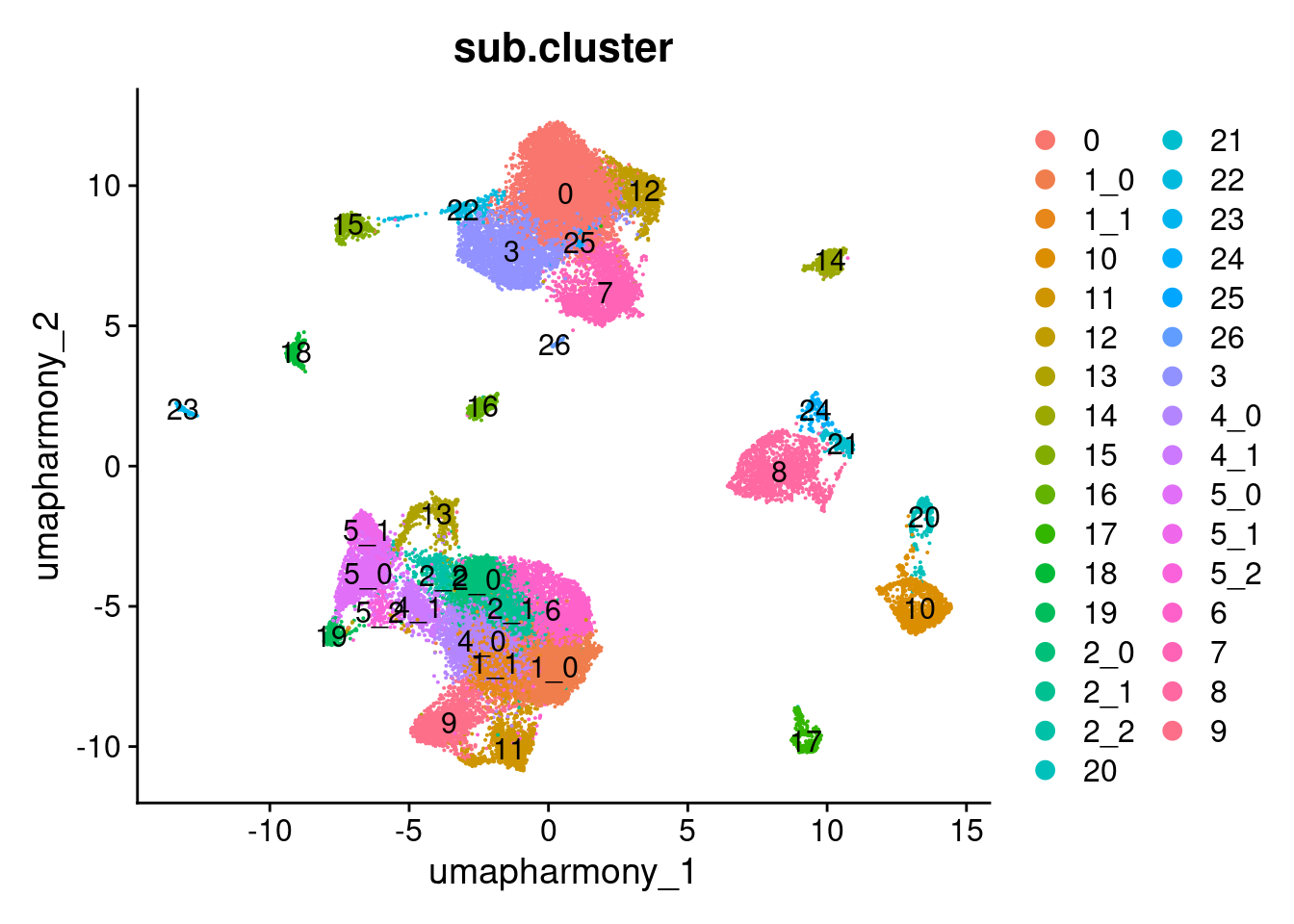
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11 12 13 14 15 16 17 18 19 2_0 2_1 2_2
5768 2588 1100 1123 1086 907 569 557 472 407 360 308 280 1464 980 407
20 21 22 23 24 25 26 3 4_0 4_1 5_0 5_1 5_2 6 7 8
262 238 228 177 175 46 33 2418 1826 537 1465 564 171 2186 1746 1483
9
1466 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13" "14" "17"
[13] "23" "1_0" "12" "4_0" "11" "9" "5_0" "6" "2_1" "2_0" "16" "5_1"
[25] "19" "2_2" "5_2" "22" "24" "15" "20" "4_1" "26" 6.4.5 Cluster 6
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "6",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 2186
Number of edges: 58961
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8570
Number of communities: 2
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)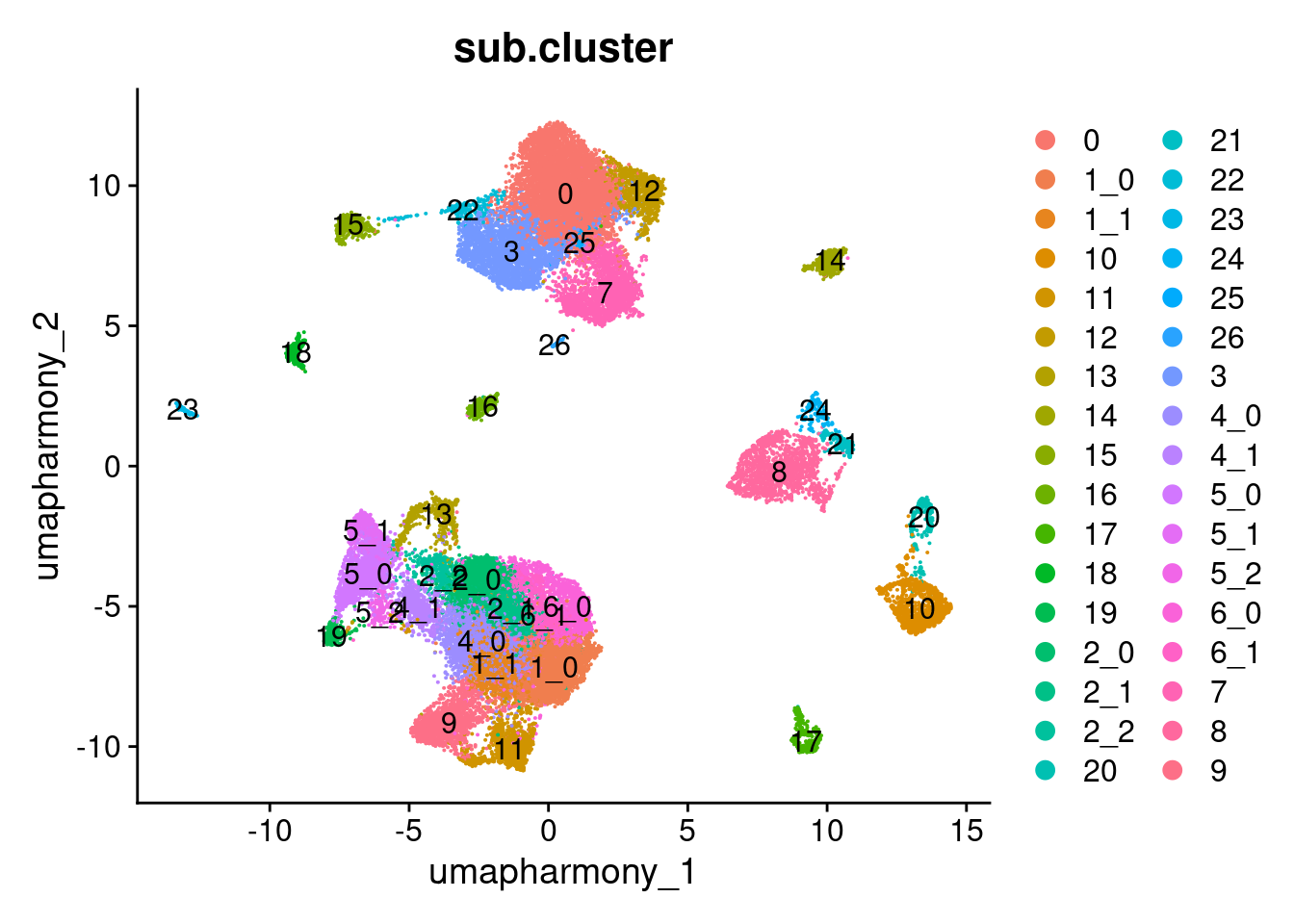
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11 12 13 14 15 16 17 18 19 2_0 2_1 2_2
5768 2588 1100 1123 1086 907 569 557 472 407 360 308 280 1464 980 407
20 21 22 23 24 25 26 3 4_0 4_1 5_0 5_1 5_2 6_0 6_1 7
262 238 228 177 175 46 33 2418 1826 537 1465 564 171 1263 923 1746
8 9
1483 1466 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13" "14" "17"
[13] "23" "1_0" "12" "4_0" "11" "9" "5_0" "6_0" "6_1" "2_1" "2_0" "16"
[25] "5_1" "19" "2_2" "5_2" "22" "24" "15" "20" "4_1" "26" 6.4.6 Cluster 9
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "9",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 1466
Number of edges: 53905
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8001
Number of communities: 2
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)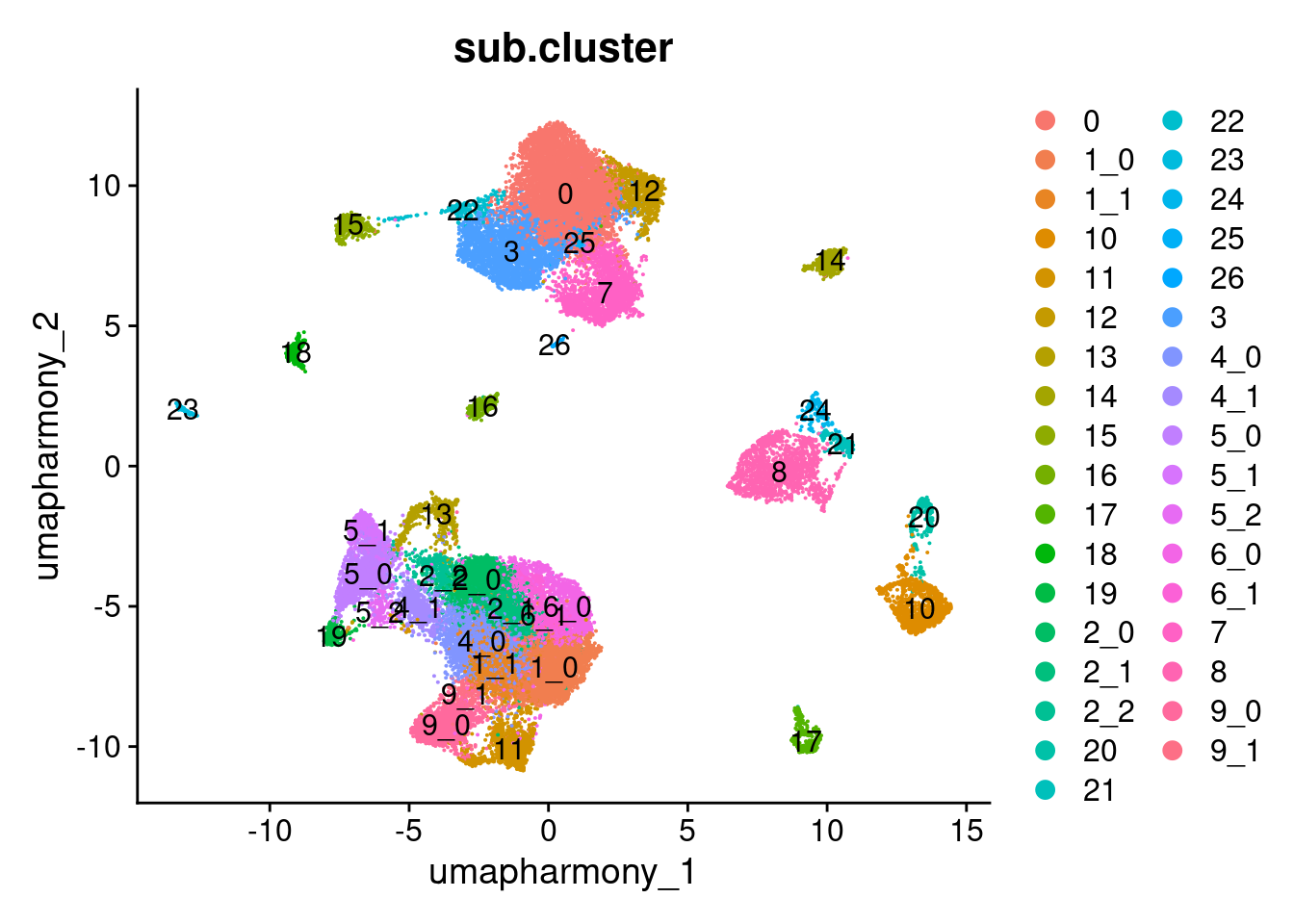
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11 12 13 14 15 16 17 18 19 2_0 2_1 2_2
5768 2588 1100 1123 1086 907 569 557 472 407 360 308 280 1464 980 407
20 21 22 23 24 25 26 3 4_0 4_1 5_0 5_1 5_2 6_0 6_1 7
262 238 228 177 175 46 33 2418 1826 537 1465 564 171 1263 923 1746
8 9_0 9_1
1483 1310 156 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13" "14" "17"
[13] "23" "1_0" "12" "4_0" "11" "9_0" "5_0" "6_0" "6_1" "2_1" "2_0" "16"
[25] "5_1" "19" "2_2" "5_2" "22" "24" "15" "20" "4_1" "9_1" "26" 6.4.7 Cluster 11
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "11",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 1086
Number of edges: 32215
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8363
Number of communities: 2
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)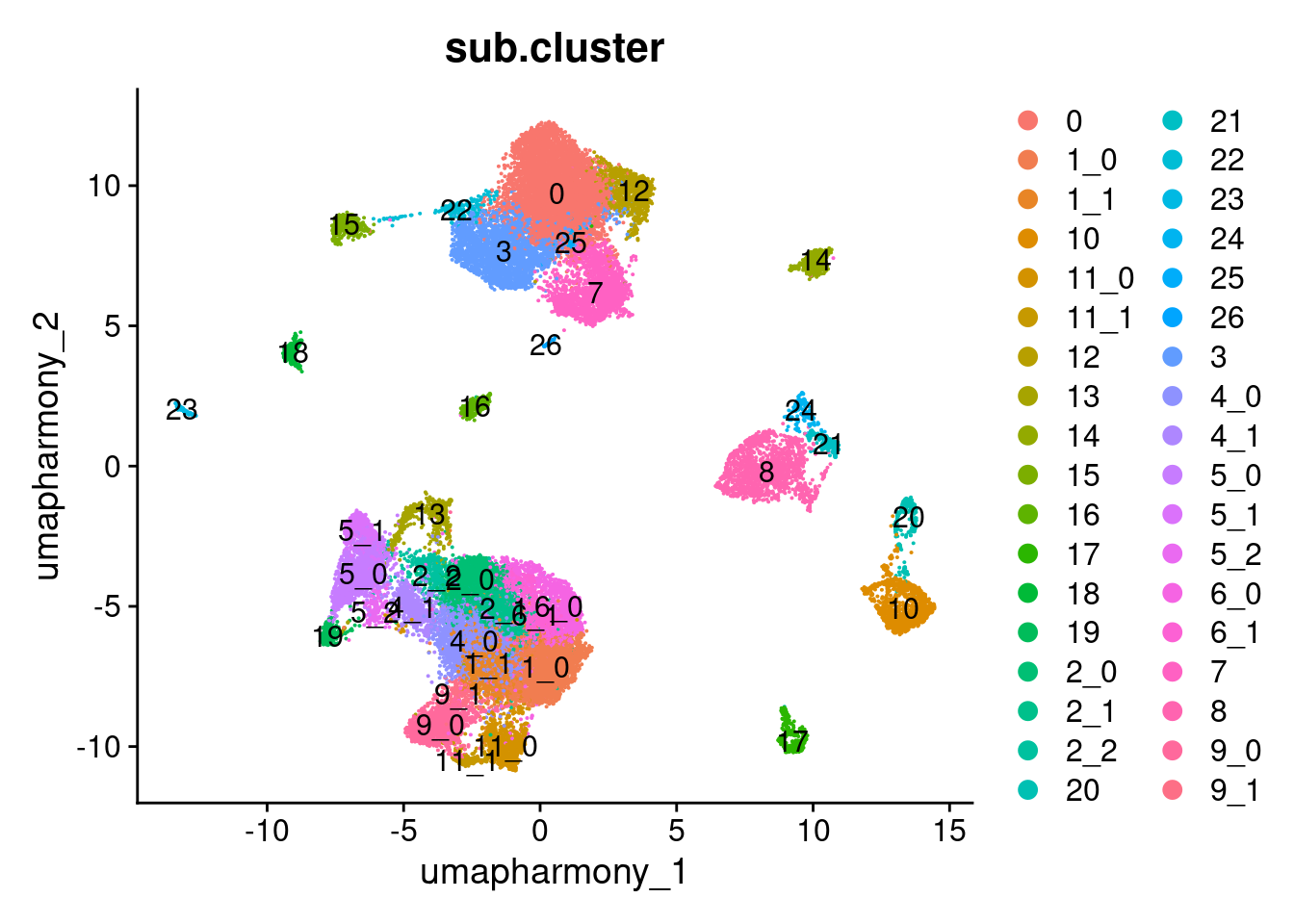
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11_0 11_1 12 13 14 15 16 17 18 19 2_0 2_1
5768 2588 1100 1123 929 157 907 569 557 472 407 360 308 280 1464 980
2_2 20 21 22 23 24 25 26 3 4_0 4_1 5_0 5_1 5_2 6_0 6_1
407 262 238 228 177 175 46 33 2418 1826 537 1465 564 171 1263 923
7 8 9_0 9_1
1746 1483 1310 156 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13"
[11] "14" "17" "23" "1_0" "12" "4_0" "11_0" "9_0" "11_1" "5_0"
[21] "6_0" "6_1" "2_1" "2_0" "16" "5_1" "19" "2_2" "5_2" "22"
[31] "24" "15" "20" "4_1" "9_1" "26" 6.4.8 Cluster 13
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "13",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 569
Number of edges: 18408
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8667
Number of communities: 2
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)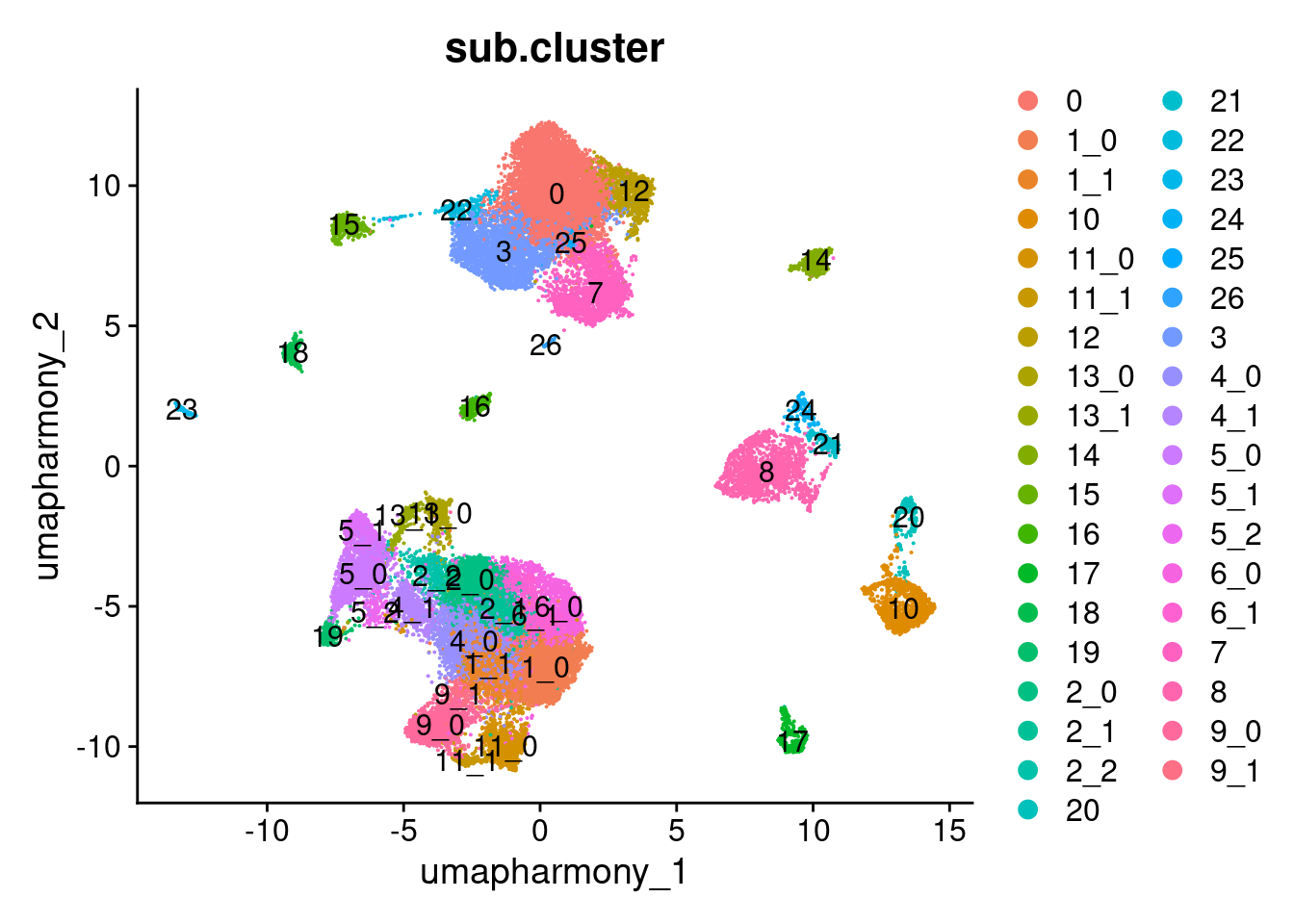
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11_0 11_1 12 13_0 13_1 14 15 16 17 18 19 2_0
5768 2588 1100 1123 929 157 907 314 255 557 472 407 360 308 280 1464
2_1 2_2 20 21 22 23 24 25 26 3 4_0 4_1 5_0 5_1 5_2 6_0
980 407 262 238 228 177 175 46 33 2418 1826 537 1465 564 171 1263
6_1 7 8 9_0 9_1
923 1746 1483 1310 156 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13_1"
[11] "14" "17" "13_0" "23" "1_0" "12" "4_0" "11_0" "9_0" "11_1"
[21] "5_0" "6_0" "6_1" "2_1" "2_0" "16" "5_1" "19" "2_2" "5_2"
[31] "22" "24" "15" "20" "4_1" "9_1" "26" 6.4.9 Cluster 19
merged.18279.skin.singlets <- FindSubCluster(object = merged.18279.skin.singlets,
cluster = "19",
resolution = 0.2,
graph.name = "RNA_snn",
algorithm = 2)Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 280
Number of edges: 10030
Running Louvain algorithm with multilevel refinement...
Maximum modularity in 10 random starts: 0.8000
Number of communities: 1
Elapsed time: 0 secondsDimPlot(merged.18279.skin.singlets,
reduction = "umap.harmony",
group.by = "sub.cluster",
label = TRUE)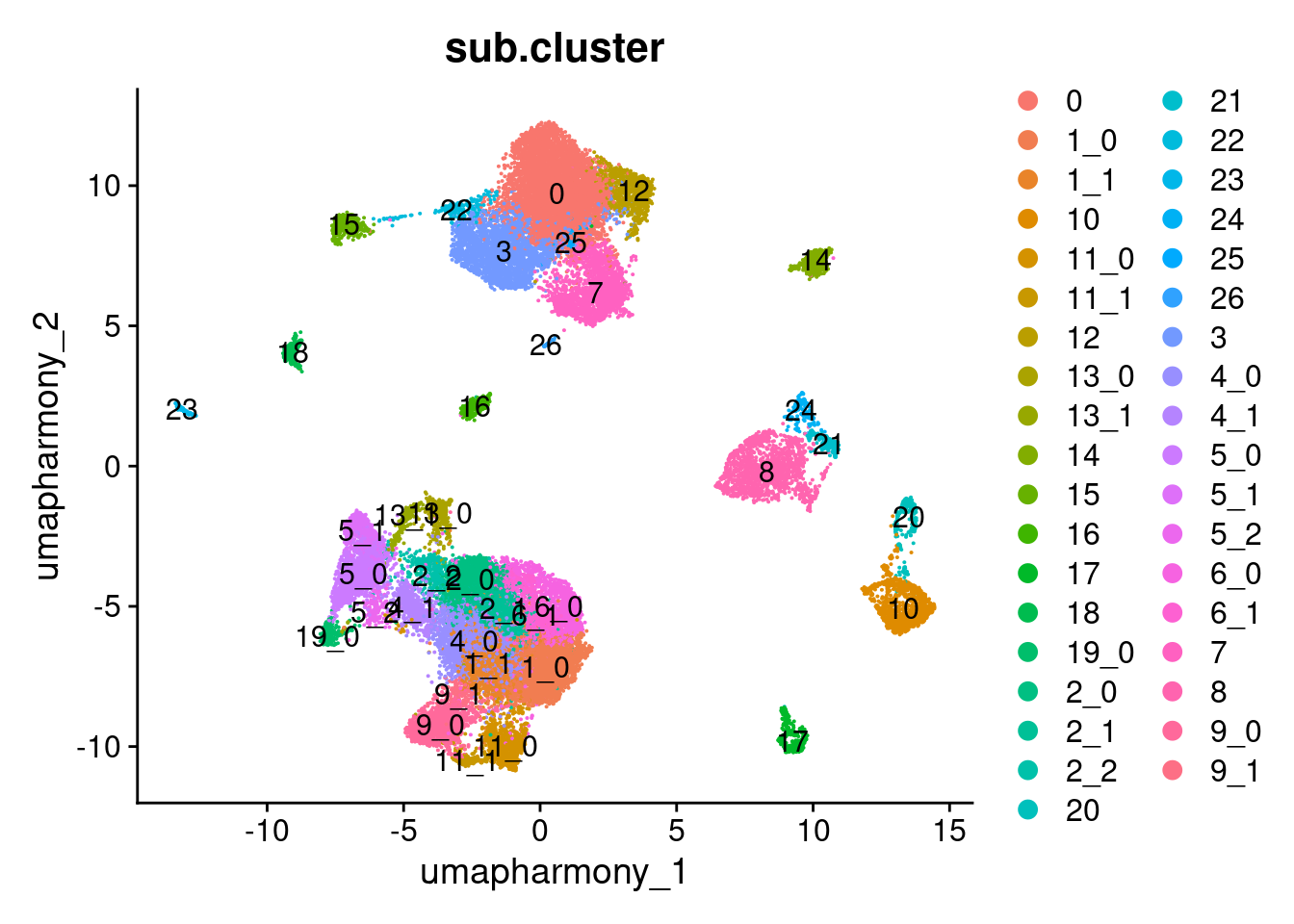
table(merged.18279.skin.singlets$sub.cluster)
0 1_0 1_1 10 11_0 11_1 12 13_0 13_1 14 15 16 17 18 19_0 2_0
5768 2588 1100 1123 929 157 907 314 255 557 472 407 360 308 280 1464
2_1 2_2 20 21 22 23 24 25 26 3 4_0 4_1 5_0 5_1 5_2 6_0
980 407 262 238 228 177 175 46 33 2418 1826 537 1465 564 171 1263
6_1 7 8 9_0 9_1
923 1746 1483 1310 156 Idents(merged.18279.skin.singlets) <- "sub.cluster"
levels(merged.18279.skin.singlets) [1] "10" "7" "0" "25" "8" "18" "21" "3" "1_1" "13_1"
[11] "14" "17" "13_0" "23" "1_0" "12" "4_0" "11_0" "9_0" "11_1"
[21] "5_0" "6_0" "6_1" "2_1" "2_0" "16" "5_1" "19_0" "2_2" "5_2"
[31] "22" "24" "15" "20" "4_1" "9_1" "26" 6.5 Plot updated QC metrics per cluster/sub-cluster
VlnPlot(merged.18279.skin.singlets, features = "nCount_RNA", group.by="sub.cluster", pt.size=0) + NoLegend()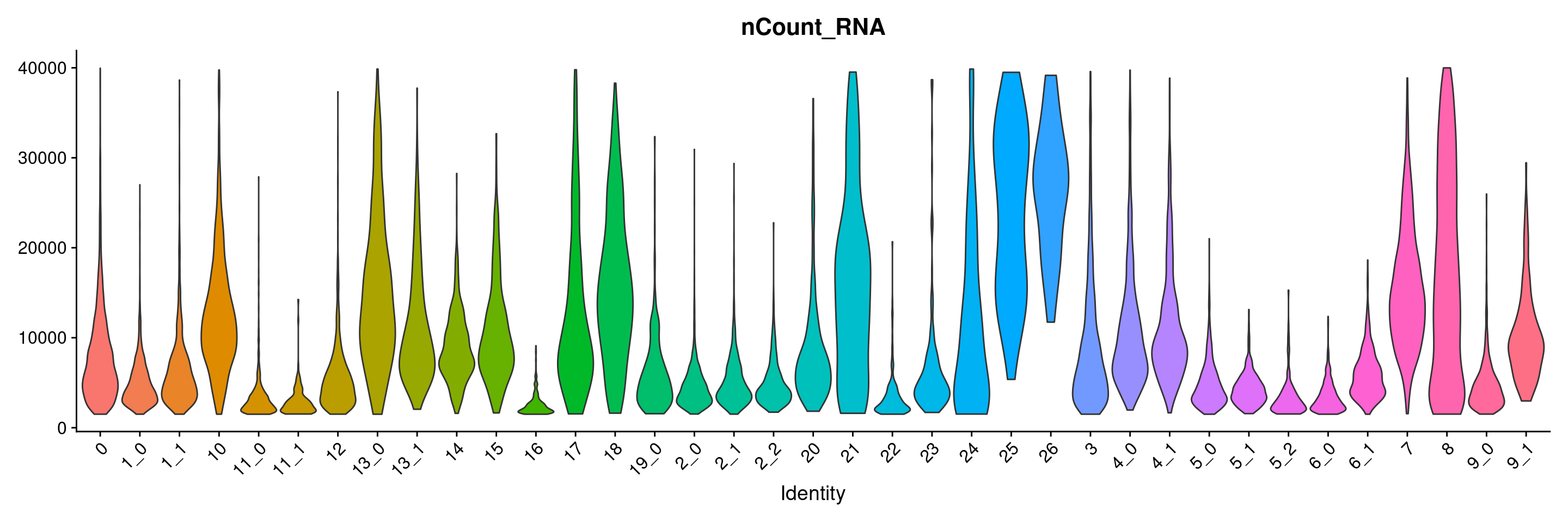
VlnPlot(merged.18279.skin.singlets, features = "nFeature_RNA", group.by="sub.cluster", pt.size=0) + NoLegend()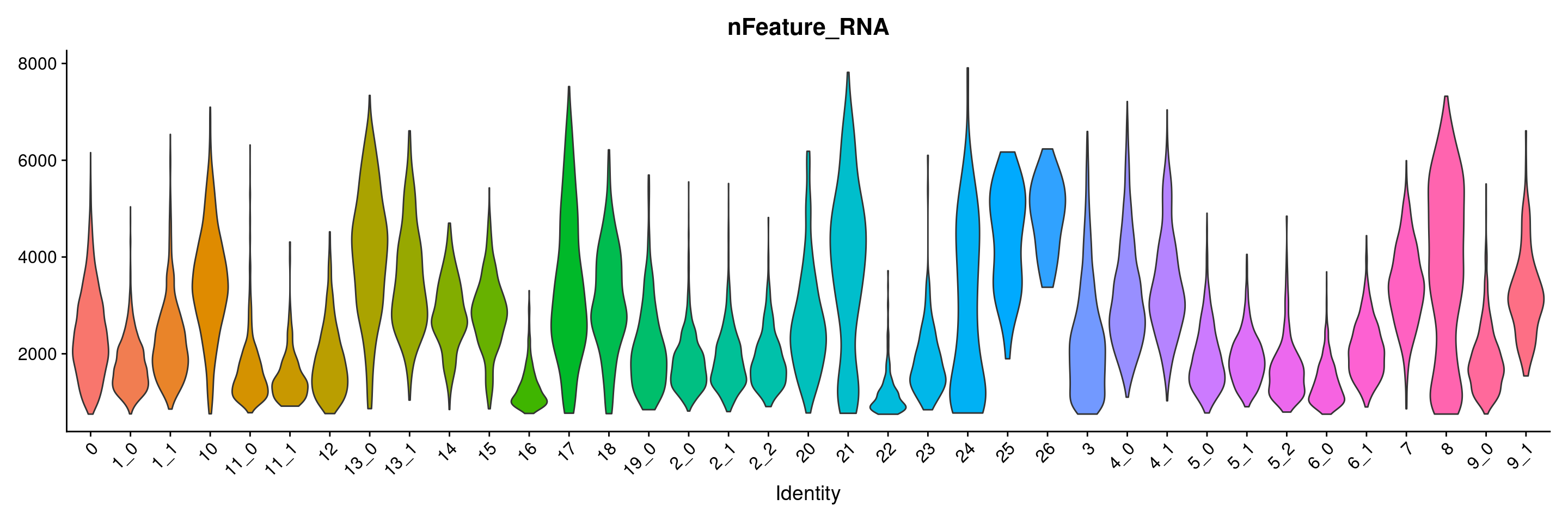
VlnPlot(merged.18279.skin.singlets, features = "percent.mt", group.by="sub.cluster", pt.size=0) + NoLegend()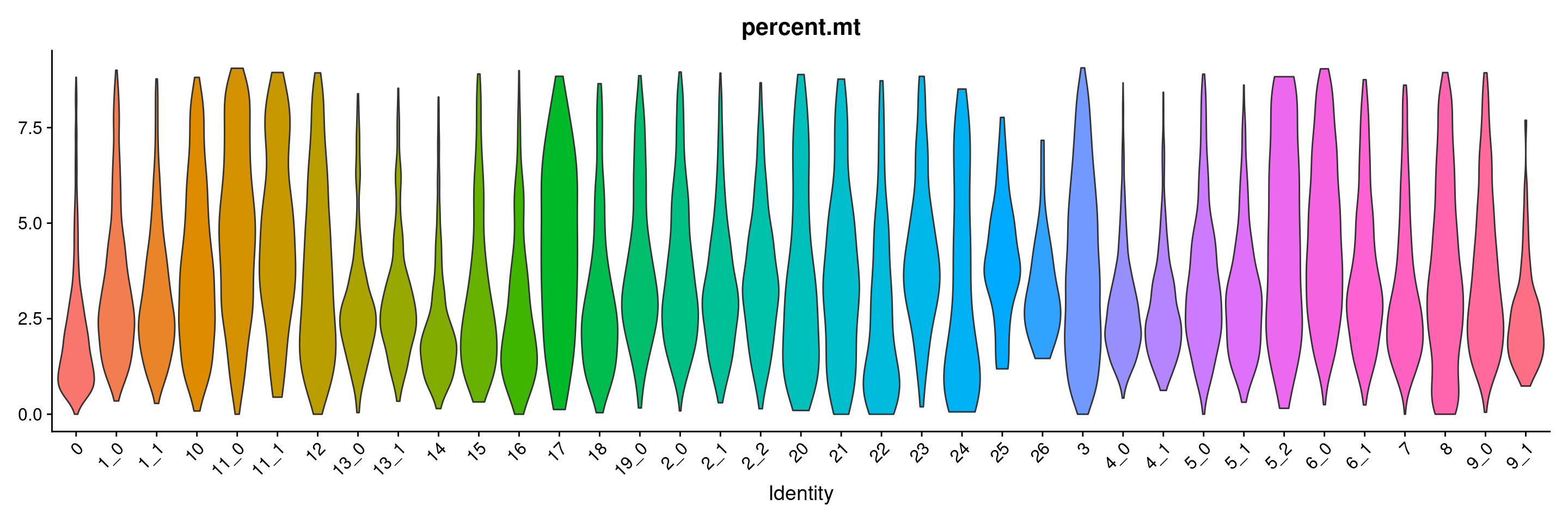
6.6 Identify updated cursory marker genes per cluster/sub-cluster
DefaultAssay(merged.18279.skin.singlets) <- "RNA"
vargenes <- presto::wilcoxauc(merged.18279.skin.singlets, 'sub.cluster', seurat_assay = 'RNA')
top_vargenes <- top_markers(vargenes, n = 100, auc_min = 0.5, pct_in_min = 50, pct_out_max = 50)
top_vargenes# A tibble: 100 × 38
rank `0` `1_0` `1_1` `10` `11_0` `11_1` `12` `13_0` `13_1` `14` `15`
<int> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
1 1 TYROBP IL7R ICOS COL6… CNOT6L LDLRA… DOCK4 STMN1 STMN1 ENSG… LAMP3
2 2 BCL2A1 LTB CD2 DCN PARP8 SKAP1 RBM47 HMGB2 TYMS IL3RA FSCN1
3 3 IL1RN SPOC… CD3E COL6… SYNE2 ZEB1 FNDC… TYMS HMGB2 IRF4 DAPP1
4 4 FCER1G CD3E CD3D COL1… BCL11B SYNE2 MYO9B RRM2 GZMB P2RY6 MARC…
5 5 IFI30 KLF2 SPOC… NNMT SLC38… STAM KYNU DUT CTSW ENSG… NUB1
6 6 MARCKS LDHB CLEC… CALD1 PRKCH CASK CYRIA MKI67 NKG7 PLD4 CERS6
7 7 PLAUR ICOS TENT… C1R PPP1R… IKZF2 ABCA1 PPP1CA GZMA SLC1… CD83
8 8 S100A9 CD6 CD6 SELE… CAMK4 CD247 DMXL2 CKS1B GNLY JCHA… CCR7
9 9 ENSG00… CD3D PDCD1 CXCL… PITPN… PRKCH ATP1… PCLAF RRM2 POLB BASP1
10 10 PLEK BCL1… BCL1… COL1… KLF12 RORA FNIP2 H4C3 PRF1 FCHS… CSF2…
# ℹ 90 more rows
# ℹ 26 more variables: `16` <chr>, `17` <chr>, `18` <chr>, `19_0` <chr>,
# `2_0` <chr>, `2_1` <chr>, `2_2` <chr>, `20` <chr>, `21` <chr>, `22` <chr>,
# `23` <chr>, `24` <chr>, `25` <chr>, `26` <chr>, `3` <chr>, `4_0` <chr>,
# `4_1` <chr>, `5_0` <chr>, `5_1` <chr>, `5_2` <chr>, `6_0` <chr>,
# `6_1` <chr>, `7` <chr>, `8` <chr>, `9_0` <chr>, `9_1` <chr>write_tsv(top_vargenes,"Skin_scRNA_prestoMarkers.tsv")6.7 Plot top markers identified and canonical genes as a dotplot
tumor <- c("DCT","MLANA","MITF","PMEL","S100A1","TYR","APOC1")
endothelial <- c("PECAM1","VWF")
fibroblast <- c("COL3A1","COL1A1","COL1A2","LUM")
tcell <- c("FGFBP2","KLRD1","CD3E","CD3D","GZMB","XCL2","GZMH","CST7","GZMK","GZMA","IFNG","GNLY","CCL4","NKG7","CCL5","CD8A","CD8B","CTLA4","TNFRSF4","BATF","ITM2A")
mono <- c("LYZ","CD74","CD68")
bcell <- c("MS4A1","CD79A")
top_vargenes <- top_markers(vargenes, n = 5, auc_min = 0.5, pct_in_min = 50, pct_out_max = 50)
top_markers <- top_vargenes %>%
select(-rank) %>%
unclass() %>%
stack() %>%
pull(values) %>%
unique() %>%
.[!is.na(.)]
dotplotmarkers <- unique(c(top_markers,tumor,endothelial,fibroblast,tcell,mono,bcell))
# Compute aggregated expression values of these genes and cluster them to order the figure
rna <- AverageExpression(merged.18279.skin.singlets,assay="RNA",slot="data")As of Seurat v5, we recommend using AggregateExpression to perform pseudo-bulk analysis.
Names of identity class contain underscores ('_'), replacing with dashes ('-')
First group.by variable `ident` starts with a number, appending `g` to ensure valid variable names
This message is displayed once per session.rna.sub <- rna$RNA[dotplotmarkers,]
cors.genes <- as.dist(1-cor(as.matrix(t(rna.sub)),method="pearson"))
hc.genes <- hclust(cors.genes)
dotplotmarkers.sorted <- rownames(rna.sub)[hc.genes$order]
# Also re-order the sub-clusters
cors.subs <- as.dist(1-cor(as.matrix(rna.sub),method="pearson"))
hc.subs <- hclust(cors.subs)
merged.18279.skin.singlets@active.ident <- factor(merged.18279.skin.singlets@active.ident,
levels=str_replace_all(str_replace_all(colnames(rna.sub)[hc.subs$order],"g",""),"-","_"))
# Plot
DotPlot(merged.18279.skin.singlets,
features = dotplotmarkers.sorted,
assay = "RNA",
cols = c("blue","red")
) +
RotatedAxis()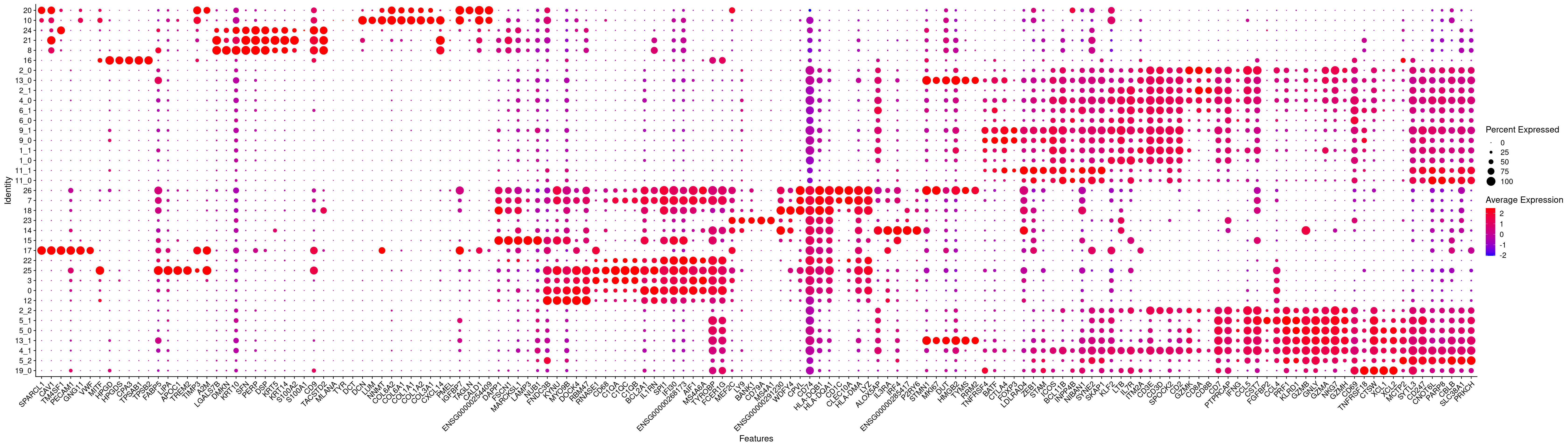
6.8 Plot updated skin sample proportions, pre vs post 3rd vaccine, per cluster
as.data.frame(table(merged.18279.skin.singlets$sub.cluster, merged.18279.skin.singlets$Sample)) %>%
as_tibble() %>%
dplyr::rename("Sample" = Var2,"Cluster" = Var1) %>%
separate(Sample, into=c("Patient","Site","Timepoint","IpiCohort","Assay"),sep="_",remove=F) %>%
dplyr::filter(Timepoint=="Pre3rd" | Timepoint=="Post3rd") %>%
group_by(Sample) %>%
mutate(Proportion = Freq / sum(Freq)) %>%
ggplot(aes(fill = Timepoint, x = fct_relevel(Timepoint,c("Pre3rd","Post3rd")), y=Proportion)) +
geom_boxplot(outlier.shape=NA) +
geom_point(pch = 21, position = position_jitterdodge(jitter.width=0.2)) +
facet_wrap(~Cluster,scales="free") +
xlab("Timepoint") +
scale_fill_discrete(breaks=c("Pre3rd","Post3rd"))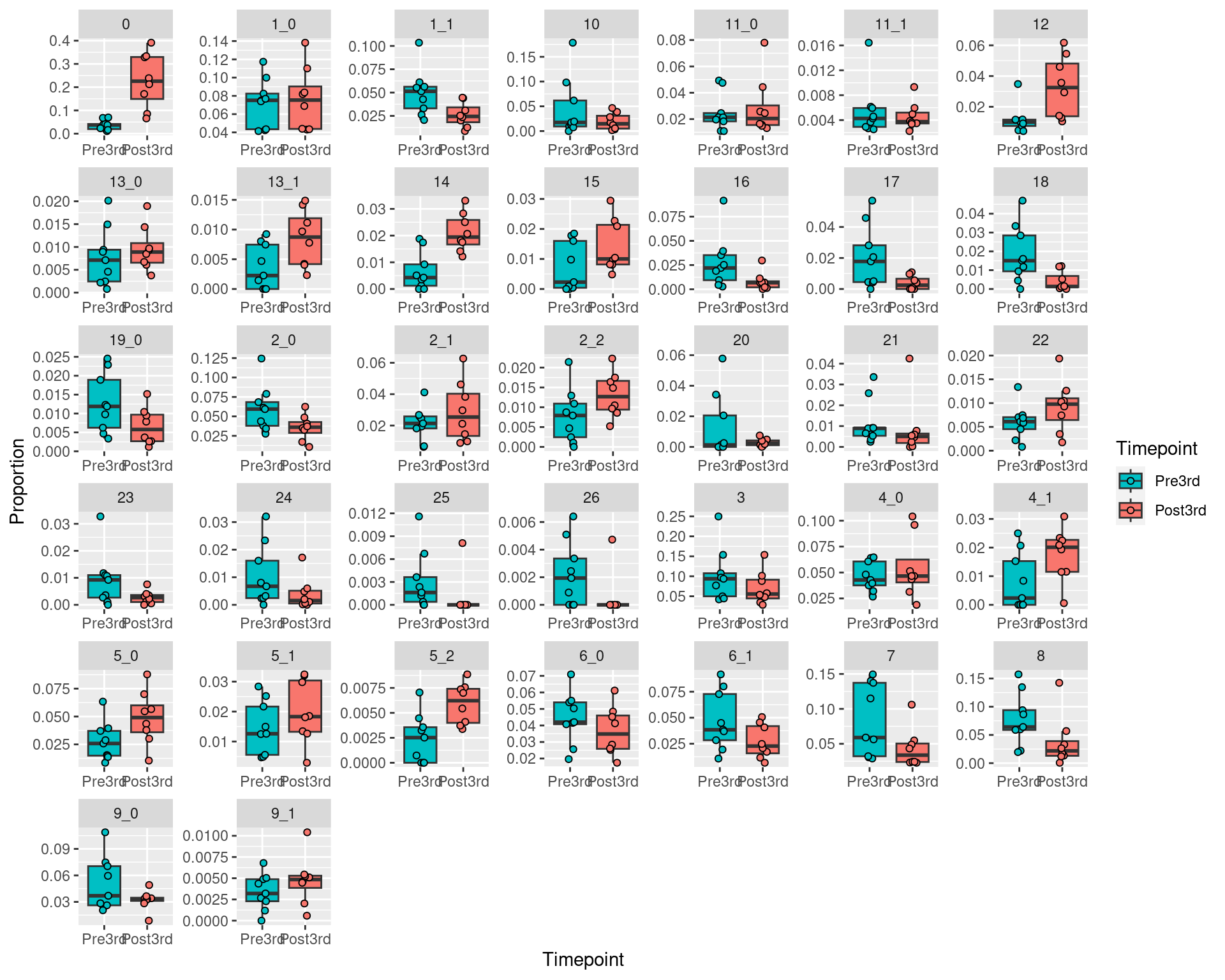
6.9 Save updated object
saveRDS(merged.18279.skin.singlets, "Skin_scRNA_Part6.rds")6.10 Get session info
sessionInfo()R version 4.3.1 (2023-06-16)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Rocky Linux 8.10 (Green Obsidian)
Matrix products: default
BLAS/LAPACK: /usr/lib64/libopenblasp-r0.3.15.so; LAPACK version 3.9.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: America/New_York
tzcode source: system (glibc)
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] cellXY_0.99.0 scDblFinder_1.14.0
[3] harmony_1.2.0 alevinQC_1.16.1
[5] vctrs_0.6.5 patchwork_1.3.0
[7] scater_1.28.0 scuttle_1.10.3
[9] speckle_1.0.0 Matrix_1.6-4
[11] fishpond_2.6.2 readxl_1.4.3
[13] SingleCellExperiment_1.22.0 SummarizedExperiment_1.30.2
[15] Biobase_2.60.0 GenomicRanges_1.52.1
[17] GenomeInfoDb_1.36.4 IRanges_2.34.1
[19] S4Vectors_0.38.2 BiocGenerics_0.46.0
[21] MatrixGenerics_1.12.3 matrixStats_1.2.0
[23] flexmix_2.3-19 lattice_0.22-5
[25] SeuratWrappers_0.3.19 miQC_1.8.0
[27] lubridate_1.9.3 forcats_1.0.0
[29] stringr_1.5.1 dplyr_1.1.4
[31] purrr_1.0.2 readr_2.1.5
[33] tidyr_1.3.1 tibble_3.2.1
[35] ggplot2_3.4.4 tidyverse_2.0.0
[37] Seurat_5.1.0 SeuratObject_5.0.2
[39] sp_2.1-3 sctransform_0.4.1
[41] glmGamPoi_1.12.2 presto_1.0.0
[43] Rcpp_1.0.12 devtools_2.4.5
[45] usethis_2.2.2 data.table_1.15.0
loaded via a namespace (and not attached):
[1] fs_1.6.3 spatstat.sparse_3.0-3
[3] bitops_1.0-7 httr_1.4.7
[5] RColorBrewer_1.1-3 profvis_0.3.8
[7] tools_4.3.1 utf8_1.2.4
[9] R6_2.5.1 DT_0.31
[11] lazyeval_0.2.2 uwot_0.1.16
[13] urlchecker_1.0.1 withr_3.0.0
[15] GGally_2.2.1 gridExtra_2.3
[17] progressr_0.14.0 cli_3.6.2
[19] spatstat.explore_3.2-6 fastDummies_1.7.3
[21] labeling_0.4.3 spatstat.data_3.0-4
[23] ggridges_0.5.6 pbapply_1.7-2
[25] Rsamtools_2.16.0 R.utils_2.12.3
[27] parallelly_1.37.0 sessioninfo_1.2.2
[29] limma_3.56.2 RSQLite_2.3.5
[31] BiocIO_1.10.0 generics_0.1.3
[33] vroom_1.6.5 gtools_3.9.5
[35] ica_1.0-3 spatstat.random_3.2-2
[37] ggbeeswarm_0.7.2 fansi_1.0.6
[39] abind_1.4-5 R.methodsS3_1.8.2
[41] lifecycle_1.0.4 yaml_2.3.8
[43] edgeR_3.42.4 Rtsne_0.17
[45] blob_1.2.4 grid_4.3.1
[47] dqrng_0.3.2 promises_1.2.1
[49] crayon_1.5.2 shinydashboard_0.7.2
[51] miniUI_0.1.1.1 beachmat_2.16.0
[53] cowplot_1.1.3 KEGGREST_1.40.1
[55] metapod_1.8.0 pillar_1.9.0
[57] knitr_1.45 rjson_0.2.21
[59] xgboost_1.7.7.1 future.apply_1.11.1
[61] codetools_0.2-19 leiden_0.4.3.1
[63] glue_1.7.0 remotes_2.4.2.1
[65] png_0.1-8 spam_2.10-0
[67] org.Mm.eg.db_3.18.0 cellranger_1.1.0
[69] gtable_0.3.4 cachem_1.0.8
[71] xfun_0.42 S4Arrays_1.2.0
[73] mime_0.12 survival_3.5-8
[75] statmod_1.5.0 bluster_1.10.0
[77] ellipsis_0.3.2 fitdistrplus_1.1-11
[79] ROCR_1.0-11 nlme_3.1-164
[81] bit64_4.0.5 RcppAnnoy_0.0.22
[83] irlba_2.3.5.1 vipor_0.4.7
[85] KernSmooth_2.23-22 DBI_1.2.2
[87] colorspace_2.1-0 nnet_7.3-19
[89] ggrastr_1.0.2 tidyselect_1.2.0
[91] bit_4.0.5 compiler_4.3.1
[93] BiocNeighbors_1.18.0 DelayedArray_0.26.7
[95] plotly_4.10.4 rtracklayer_1.60.1
[97] scales_1.3.0 lmtest_0.9-40
[99] digest_0.6.34 goftest_1.2-3
[101] spatstat.utils_3.0-4 rmarkdown_2.25
[103] XVector_0.40.0 htmltools_0.5.7
[105] pkgconfig_2.0.3 sparseMatrixStats_1.12.2
[107] fastmap_1.1.1 rlang_1.1.3
[109] htmlwidgets_1.6.4 shiny_1.8.0
[111] DelayedMatrixStats_1.22.6 farver_2.1.1
[113] zoo_1.8-12 jsonlite_1.8.8
[115] BiocParallel_1.34.2 R.oo_1.26.0
[117] BiocSingular_1.16.0 RCurl_1.98-1.14
[119] magrittr_2.0.3 modeltools_0.2-23
[121] GenomeInfoDbData_1.2.10 dotCall64_1.1-1
[123] munsell_0.5.0 viridis_0.6.5
[125] reticulate_1.35.0 stringi_1.8.3
[127] zlibbioc_1.46.0 MASS_7.3-60.0.1
[129] org.Hs.eg.db_3.18.0 plyr_1.8.9
[131] pkgbuild_1.4.3 ggstats_0.5.1
[133] parallel_4.3.1 listenv_0.9.1
[135] ggrepel_0.9.5 deldir_2.0-2
[137] Biostrings_2.68.1 splines_4.3.1
[139] tensor_1.5 hms_1.1.3
[141] locfit_1.5-9.8 igraph_2.0.2
[143] spatstat.geom_3.2-8 RcppHNSW_0.6.0
[145] reshape2_1.4.4 ScaledMatrix_1.8.1
[147] pkgload_1.3.4 XML_3.99-0.16.1
[149] evaluate_0.23 scran_1.28.2
[151] BiocManager_1.30.22 tzdb_0.4.0
[153] httpuv_1.6.14 RANN_2.6.1
[155] polyclip_1.10-6 future_1.33.1
[157] scattermore_1.2 rsvd_1.0.5
[159] xtable_1.8-4 restfulr_0.0.15
[161] svMisc_1.2.3 RSpectra_0.16-1
[163] later_1.3.2 viridisLite_0.4.2
[165] AnnotationDbi_1.64.1 GenomicAlignments_1.36.0
[167] memoise_2.0.1 beeswarm_0.4.0
[169] tximport_1.28.0 cluster_2.1.6
[171] timechange_0.3.0 globals_0.16.2