# Load libraries
library(data.table)
library(devtools)
library(presto)
library(glmGamPoi)
library(sctransform)
library(Seurat)
library(tidyverse)
library(miQC)
library(SeuratWrappers)
library(flexmix)
library(SingleCellExperiment)
library(SummarizedExperiment)
library(readxl)
library(fishpond)
library(Matrix)
library(speckle)
library(scater)
library(patchwork)
library(vctrs)
library(alevinQC)
library(harmony)
library(scDblFinder)
library(cellXY)
library(STACAS)
library(ProjecTILs)
# Set global options for Seurat v5 objects
options(Seurat.object.assay.version = 'v5')8 Skin: Classification of CD4 and CD8 T cell states within T/NK lineage
8.1 Set up Seurat workspace
8.2 Load previously saved sub-clustered object
merged.18279.skin.singlets <- readRDS("Skin_scRNA_Part6.rds")8.3 Run CD8 and CD4 projection with projecTILs as in this vignette
Note these classifications are limited only to CD4/CD8 and do not include NK cells
8.3.1 Retrieve human CD8 and CD4 reference maps
options(timeout = max(900, getOption("timeout")))
download.file("https://figshare.com/ndownloader/files/41414556", destfile = "CD8T_human_ref_v1.rds")
download.file("https://figshare.com/ndownloader/files/43794750", destfile = "CD4T_human_ref_v2.rds")8.3.2 Load reference maps
ref.cd8 <- load.reference.map("CD8T_human_ref_v1.rds")[1] "Loading Custom Reference Atlas..."
[1] "Loaded Custom Reference map Human CD8 TILs"ref.cd4 <- load.reference.map("CD4T_human_ref_v2.rds")[1] "Loading Custom Reference Atlas..."
[1] "Loaded Custom Reference map custom_reference"8.3.3 Plot reference atlases
a <- DimPlot(ref.cd8, cols = ref.cd8@misc$atlas.palette, label = T) + theme(aspect.ratio = 1) +
ggtitle("CD8 T reference") + NoLegend()
b <- DimPlot(ref.cd4, cols = ref.cd4@misc$atlas.palette, label = T) + theme(aspect.ratio = 1) +
ggtitle("CD4 T reference") + NoLegend()
a | b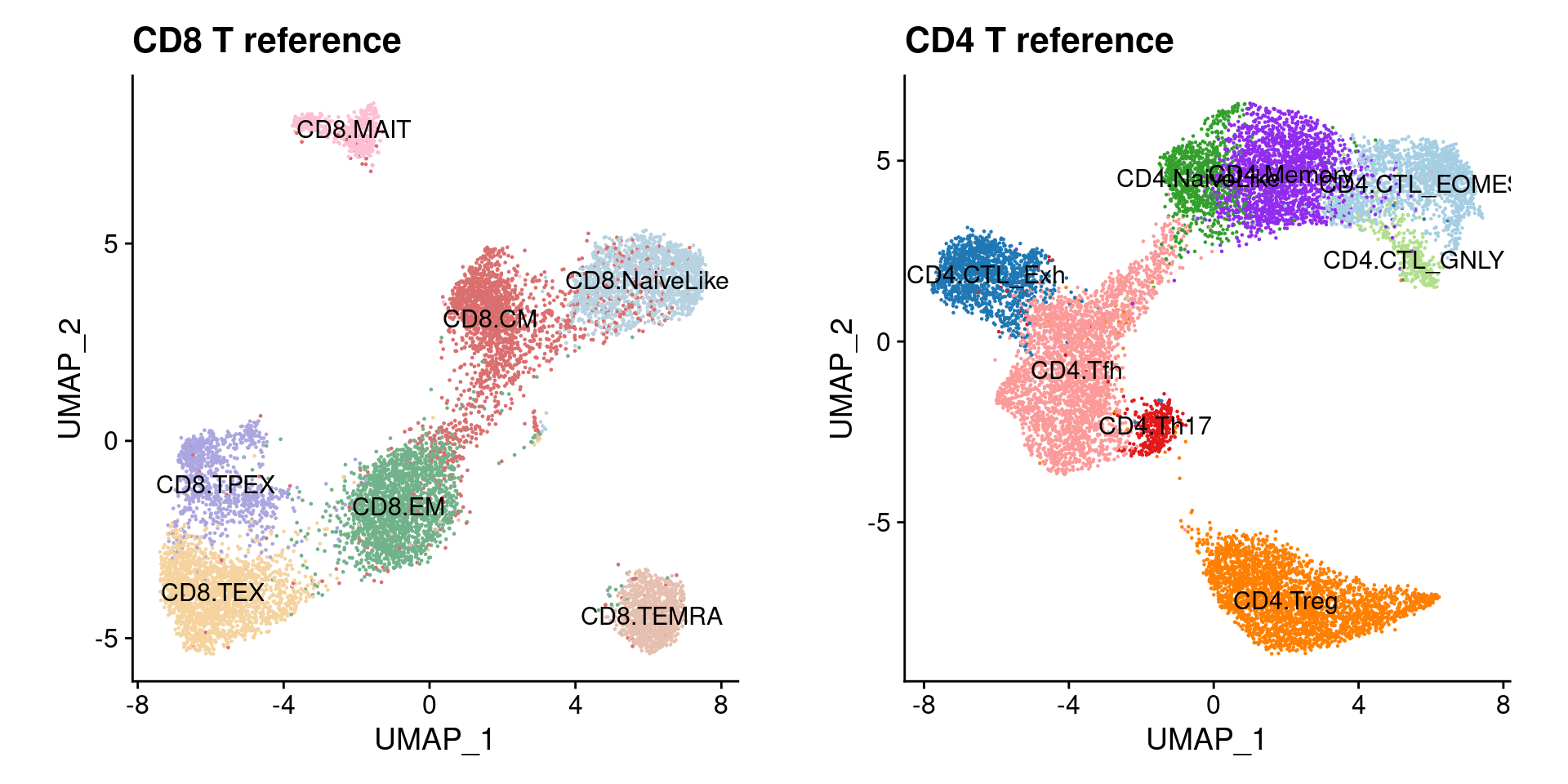
8.3.4 Classify CD8 T subtypes
merged.18279.skin.singlets <- ProjecTILs.classifier(merged.18279.skin.singlets, ref.cd8, ncores = 8, split.by = "Sample")
|
| | 0%[1] "Using assay RNA for query"
[1] "383 out of 437 ( 88% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1255 out of 1342 ( 94% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "339 out of 426 ( 80% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1501 out of 1718 ( 87% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "743 out of 818 ( 91% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1221 out of 1375 ( 89% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "974 out of 1191 ( 82% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1284 out of 1481 ( 87% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1516 out of 1655 ( 92% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "881 out of 937 ( 94% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "2364 out of 2689 ( 88% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "2658 out of 3429 ( 78% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "2212 out of 2526 ( 88% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1008 out of 1121 ( 90% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "3610 out of 3726 ( 97% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "4680 out of 5434 ( 86% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "2661 out of 3092 ( 86% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%8.3.5 Classify CD4 T subtypes
merged.18279.skin.singlets <- ProjecTILs.classifier(merged.18279.skin.singlets, ref.cd4, ncores = 8, split.by = "Sample", overwrite = F)
|
| | 0%[1] "Using assay RNA for query"
[1] "361 out of 437 ( 83% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1139 out of 1342 ( 85% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "325 out of 426 ( 76% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1519 out of 1718 ( 88% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "483 out of 818 ( 59% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1102 out of 1375 ( 80% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "849 out of 1191 ( 71% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1101 out of 1481 ( 74% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1363 out of 1655 ( 82% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "731 out of 937 ( 78% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "1854 out of 2689 ( 69% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "2485 out of 3429 ( 72% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "2158 out of 2526 ( 85% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "819 out of 1121 ( 73% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "3262 out of 3726 ( 88% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "4574 out of 5434 ( 84% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%
|
| | 0%[1] "Using assay RNA for query"
[1] "2233 out of 3092 ( 72% ) non-pure cells removed. Use filter.cells=FALSE to avoid pre-filtering"
Performing log-normalization
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
**************************************************|
[1] "Aligning query to reference map for batch-correction..."
Projecting corrected query onto Reference PCA space
Projecting corrected query onto Reference UMAP space
|
|======================================================================| 100%8.4 Tabulate and visualize projections
In addition to guiding annotation, these projected cell states can be quantified downstream comparing pre vs post-vaccine, tumor samples over time, or to characterize the phenotypes of T-cells that have expanded with vaccination
8.4.1 Count total cells by T cell state
table(merged.18279.skin.singlets$functional.cluster, useNA = "ifany")
CD4.CTL_EOMES CD4.CTL_Exh CD4.CTL_GNLY CD4.Memory CD4.NaiveLike
1110 166 534 1742 1486
CD4.Tfh CD4.Th17 CD4.Treg CD8.CM CD8.EM
575 84 1329 2312 1092
CD8.MAIT CD8.NaiveLike CD8.TEMRA CD8.TEX CD8.TPEX
47 239 77 266 61
<NA>
22277 8.4.2 Tabulate which T cell state was most abundant in each subcluster
as.data.frame(table(merged.18279.skin.singlets$sub.cluster, merged.18279.skin.singlets$functional.cluster, useNA = "ifany")) %>%
as_tibble() %>%
dplyr::rename("Cluster" = Var1, "CellState" = Var2) %>%
group_by(Cluster) %>%
slice_max(Freq,n=1) %>%
inner_join(enframe(table(merged.18279.skin.singlets$sub.cluster),name="Cluster",value="TotalCount"),by="Cluster") %>%
as.data.frame() Cluster CellState Freq TotalCount
1 0 <NA> 5768 5768
2 1_0 CD4.NaiveLike 1093 2588
3 1_1 CD4.CTL_EOMES 454 1100
4 10 <NA> 1123 1123
5 11_0 <NA> 660 929
6 11_1 <NA> 106 157
7 12 <NA> 907 907
8 13_0 CD8.TEX 67 314
9 13_1 <NA> 247 255
10 14 <NA> 557 557
11 15 <NA> 472 472
12 16 <NA> 407 407
13 17 <NA> 360 360
14 18 <NA> 308 308
15 19_0 <NA> 274 280
16 2_0 CD8.EM 766 1464
17 2_1 CD8.CM 696 980
18 2_2 <NA> 142 407
19 20 <NA> 262 262
20 21 <NA> 238 238
21 22 <NA> 228 228
22 23 <NA> 175 177
23 24 <NA> 175 175
24 25 <NA> 46 46
25 26 <NA> 33 33
26 3 <NA> 2418 2418
27 4_0 CD8.CM 664 1826
28 4_1 <NA> 343 537
29 5_0 <NA> 1421 1465
30 5_1 <NA> 562 564
31 5_2 <NA> 169 171
32 6_0 CD4.Memory 446 1263
33 6_1 CD4.Memory 301 923
34 7 <NA> 1746 1746
35 8 <NA> 1483 1483
36 9_0 CD4.Treg 1092 1310
37 9_1 CD4.Treg 82 1568.4.3 Plot UMAP of all cells labeled by projected T cell state
DimPlot(merged.18279.skin.singlets, group.by = "functional.cluster", reduction="umap.harmony")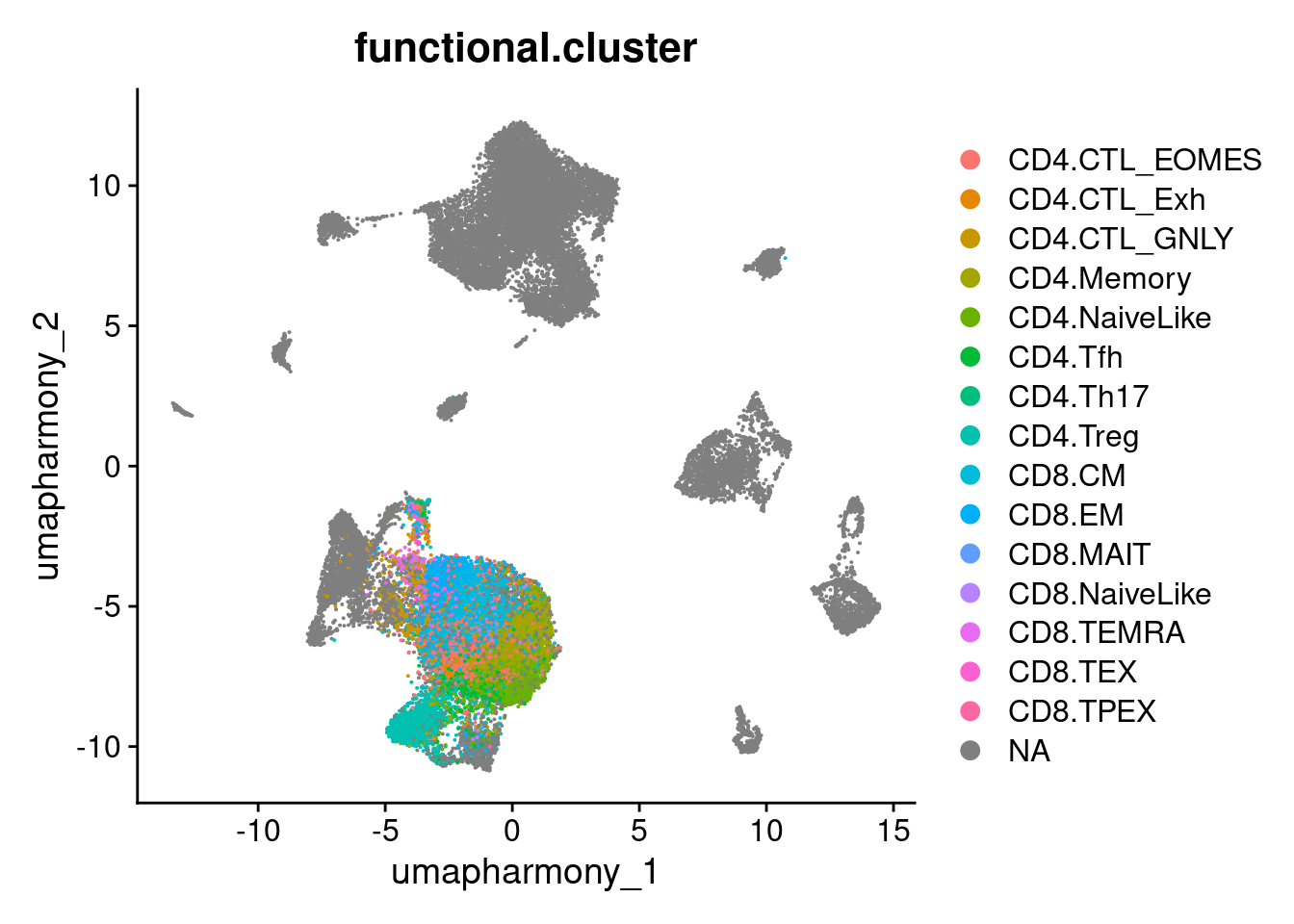
8.4.4 Plot just T/NK cell lineage
tnk_clusters <- sort(str_subset(unique(merged.18279.skin.singlets$sub.cluster),"_"))
tnk_clusters [1] "1_0" "1_1" "11_0" "11_1" "13_0" "13_1" "19_0" "2_0" "2_1" "2_2"
[11] "4_0" "4_1" "5_0" "5_1" "5_2" "6_0" "6_1" "9_0" "9_1" tnk_cells <- WhichCells(merged.18279.skin.singlets,idents = tnk_clusters)
DimPlot(merged.18279.skin.singlets,
group.by = "functional.cluster",
reduction="umap.harmony",
cells = tnk_cells)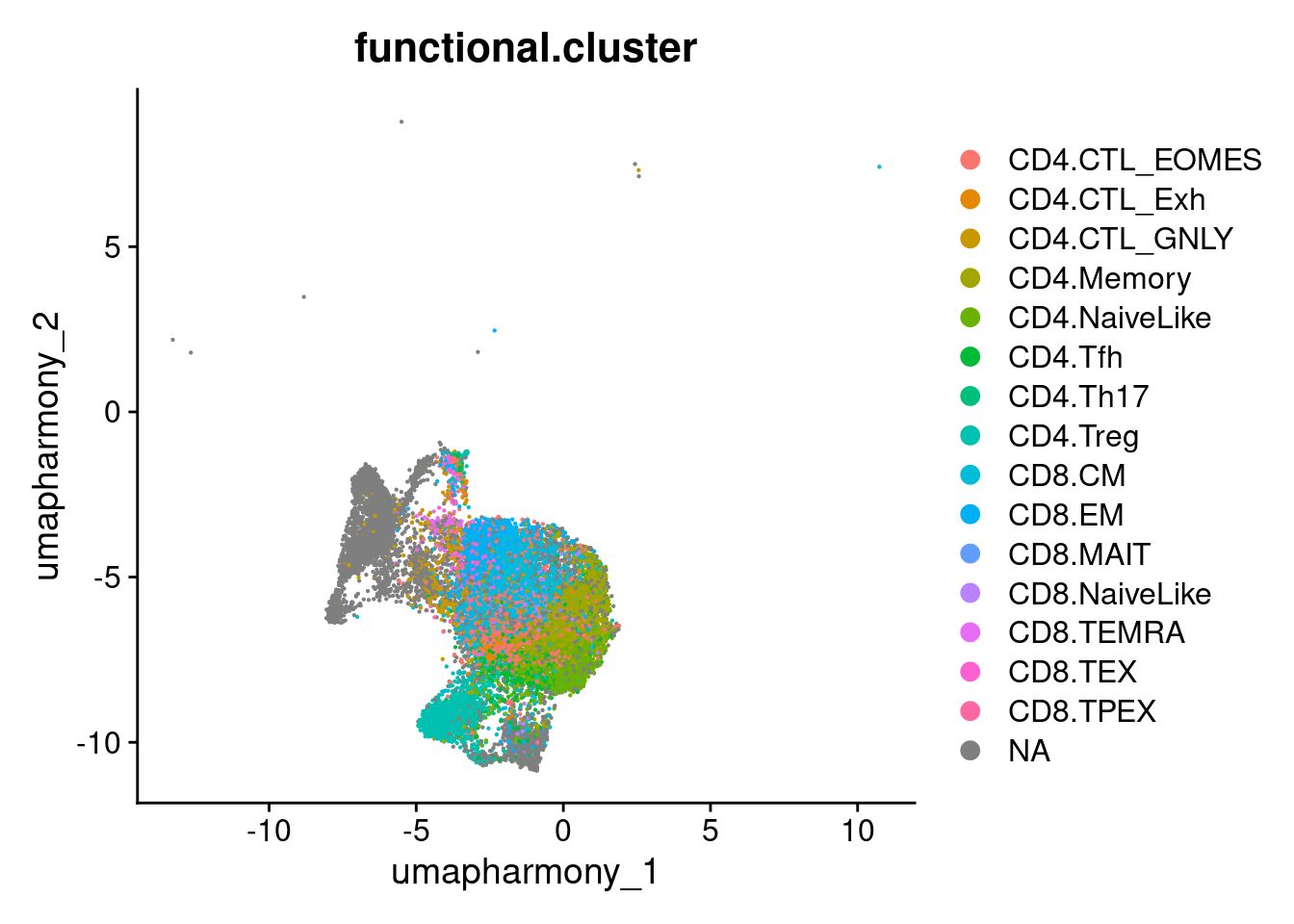
8.5 Save modified object
saveRDS(merged.18279.skin.singlets, "Skin_scRNA_Part8.rds")8.6 Get session info
sessionInfo()R version 4.3.1 (2023-06-16)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Rocky Linux 8.10 (Green Obsidian)
Matrix products: default
BLAS/LAPACK: /usr/lib64/libopenblasp-r0.3.15.so; LAPACK version 3.9.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: America/New_York
tzcode source: system (glibc)
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] ProjecTILs_3.3.1 STACAS_2.2.2
[3] cellXY_0.99.0 scDblFinder_1.14.0
[5] harmony_1.2.0 alevinQC_1.16.1
[7] vctrs_0.6.5 patchwork_1.3.0
[9] scater_1.28.0 scuttle_1.10.3
[11] speckle_1.0.0 Matrix_1.6-4
[13] fishpond_2.6.2 readxl_1.4.3
[15] SingleCellExperiment_1.22.0 SummarizedExperiment_1.30.2
[17] Biobase_2.60.0 GenomicRanges_1.52.1
[19] GenomeInfoDb_1.36.4 IRanges_2.34.1
[21] S4Vectors_0.38.2 BiocGenerics_0.46.0
[23] MatrixGenerics_1.12.3 matrixStats_1.2.0
[25] flexmix_2.3-19 lattice_0.22-5
[27] SeuratWrappers_0.3.19 miQC_1.8.0
[29] lubridate_1.9.3 forcats_1.0.0
[31] stringr_1.5.1 dplyr_1.1.4
[33] purrr_1.0.2 readr_2.1.5
[35] tidyr_1.3.1 tibble_3.2.1
[37] ggplot2_3.4.4 tidyverse_2.0.0
[39] Seurat_5.1.0 SeuratObject_5.0.2
[41] sp_2.1-3 sctransform_0.4.1
[43] glmGamPoi_1.12.2 presto_1.0.0
[45] Rcpp_1.0.12 devtools_2.4.5
[47] usethis_2.2.2 data.table_1.15.0
loaded via a namespace (and not attached):
[1] fs_1.6.3 spatstat.sparse_3.0-3
[3] bitops_1.0-7 httr_1.4.7
[5] RColorBrewer_1.1-3 profvis_0.3.8
[7] tools_4.3.1 utf8_1.2.4
[9] R6_2.5.1 DT_0.31
[11] lazyeval_0.2.2 uwot_0.1.16
[13] urlchecker_1.0.1 withr_3.0.0
[15] GGally_2.2.1 gridExtra_2.3
[17] progressr_0.14.0 cli_3.6.2
[19] spatstat.explore_3.2-6 fastDummies_1.7.3
[21] labeling_0.4.3 spatstat.data_3.0-4
[23] askpass_1.2.0 ggridges_0.5.6
[25] pbapply_1.7-2 Rsamtools_2.16.0
[27] R.utils_2.12.3 parallelly_1.37.0
[29] sessioninfo_1.2.2 limma_3.56.2
[31] RSQLite_2.3.5 BiocIO_1.10.0
[33] generics_0.1.3 gtools_3.9.5
[35] ica_1.0-3 spatstat.random_3.2-2
[37] ggbeeswarm_0.7.2 fansi_1.0.6
[39] abind_1.4-5 R.methodsS3_1.8.2
[41] lifecycle_1.0.4 yaml_2.3.8
[43] edgeR_3.42.4 Rtsne_0.17
[45] blob_1.2.4 grid_4.3.1
[47] dqrng_0.3.2 promises_1.2.1
[49] crayon_1.5.2 shinydashboard_0.7.2
[51] miniUI_0.1.1.1 beachmat_2.16.0
[53] cowplot_1.1.3 KEGGREST_1.40.1
[55] scGate_1.6.0 metapod_1.8.0
[57] pillar_1.9.0 knitr_1.45
[59] rjson_0.2.21 xgboost_1.7.7.1
[61] future.apply_1.11.1 codetools_0.2-19
[63] leiden_0.4.3.1 glue_1.7.0
[65] remotes_2.4.2.1 png_0.1-8
[67] spam_2.10-0 org.Mm.eg.db_3.18.0
[69] cellranger_1.1.0 gtable_0.3.4
[71] cachem_1.0.8 xfun_0.42
[73] S4Arrays_1.2.0 mime_0.12
[75] pracma_2.4.4 survival_3.5-8
[77] pheatmap_1.0.12 statmod_1.5.0
[79] bluster_1.10.0 ellipsis_0.3.2
[81] fitdistrplus_1.1-11 ROCR_1.0-11
[83] nlme_3.1-164 bit64_4.0.5
[85] RcppAnnoy_0.0.22 irlba_2.3.5.1
[87] vipor_0.4.7 KernSmooth_2.23-22
[89] DBI_1.2.2 colorspace_2.1-0
[91] nnet_7.3-19 UCell_2.6.2
[93] tidyselect_1.2.0 bit_4.0.5
[95] compiler_4.3.1 BiocNeighbors_1.18.0
[97] DelayedArray_0.26.7 plotly_4.10.4
[99] rtracklayer_1.60.1 scales_1.3.0
[101] lmtest_0.9-40 digest_0.6.34
[103] goftest_1.2-3 spatstat.utils_3.0-4
[105] rmarkdown_2.25 XVector_0.40.0
[107] htmltools_0.5.7 pkgconfig_2.0.3
[109] umap_0.2.10.0 sparseMatrixStats_1.12.2
[111] fastmap_1.1.1 rlang_1.1.3
[113] htmlwidgets_1.6.4 shiny_1.8.0
[115] DelayedMatrixStats_1.22.6 farver_2.1.1
[117] zoo_1.8-12 jsonlite_1.8.8
[119] BiocParallel_1.34.2 R.oo_1.26.0
[121] BiocSingular_1.16.0 RCurl_1.98-1.14
[123] magrittr_2.0.3 modeltools_0.2-23
[125] GenomeInfoDbData_1.2.10 dotCall64_1.1-1
[127] munsell_0.5.0 viridis_0.6.5
[129] reticulate_1.35.0 stringi_1.8.3
[131] zlibbioc_1.46.0 MASS_7.3-60.0.1
[133] org.Hs.eg.db_3.18.0 plyr_1.8.9
[135] pkgbuild_1.4.3 ggstats_0.5.1
[137] parallel_4.3.1 listenv_0.9.1
[139] ggrepel_0.9.5 deldir_2.0-2
[141] Biostrings_2.68.1 splines_4.3.1
[143] tensor_1.5 hms_1.1.3
[145] locfit_1.5-9.8 igraph_2.0.2
[147] spatstat.geom_3.2-8 RcppHNSW_0.6.0
[149] reshape2_1.4.4 ScaledMatrix_1.8.1
[151] pkgload_1.3.4 XML_3.99-0.16.1
[153] evaluate_0.23 scran_1.28.2
[155] BiocManager_1.30.22 tzdb_0.4.0
[157] httpuv_1.6.14 openssl_2.1.1
[159] RANN_2.6.1 polyclip_1.10-6
[161] future_1.33.1 scattermore_1.2
[163] rsvd_1.0.5 xtable_1.8-4
[165] restfulr_0.0.15 svMisc_1.2.3
[167] RSpectra_0.16-1 later_1.3.2
[169] viridisLite_0.4.2 AnnotationDbi_1.64.1
[171] GenomicAlignments_1.36.0 memoise_2.0.1
[173] beeswarm_0.4.0 tximport_1.28.0
[175] cluster_2.1.6 timechange_0.3.0
[177] globals_0.16.2